surfaceome treemap
- enzymes
- transporters
- receptors
- miscellaneous
- unclassified
- unmatched
downloads
The following items are available for download:
how to cite
The in silico human surfaceome has been published in PNAS:
The in silico human surfaceome.
Bausch-Fluck D, Goldmann U, Müller S, van Oostrum M, Müller M, Schubert OT, Wollscheid B.
Proc Natl Acad Sci U S A. 2018 Nov 13;115(46):E10988-E10997.
figures
Figure 1
Surfaceome definition and construction. (A) Visual representation of surfaceome definition. Proteins shown in red are regarded as surfaceome members; those in blue are not. (B) Compositions of nonsurface (negative) and surface (positive) training sets used for the machine-learning model. Subcellular location of the nonsurface training set are labeled as follows: 1, endoplasmic reticulum; 2, endosome; 3, Golgi apparatus; 4, lysosome; 5, mitochondrion; 6, nucleus; 7, peroxisome; 8, cytosol; and 9, multiple locations. (C) Receiver operating characteristics for the full model derived from out-of-bag error estimates (red line) (Dataset S1, 11.6). Gray line indicates the performance of random guessing. The three SURFY score cutoffs at 1%, 5%, and 15% FPRs are indicated. (D) Distribution of the predicted scores for the training sets (Upper) and for the remaining ɑ-helical TM proteins (Lower). The bars of the nonsurface training set are stacked on top of the bars of the surface training set. Score cutoffs for estimated 1%, 5%, and 15% FPRs are indicated at the bottom. The predicted score distribution for CD antigens in and outside the training set are highlighted in yellow. (E) Gini index scores (41) for the 10 most important features used in building the predictive random forest model used by SURFY. Scores are plotted as means ± SDs. Features used for calculating SURFY scores are highlighted in red. AUC, area under the curve; Avg., average; C-glyc., C-glycosylation; TMD, TM domain.
Figure 2
Characterization of the predicted surfaceome. (A–C) Comparison of 2,886 surfaceome proteins in red, with 2,216 nonsurfaceome membrane proteins in blue. (A) Distributions of sequence features in noncytoplasmic domains of surfaceome (upper graphs) and nonsurfaceome membrane (lower graphs) proteins, calculated as frequency per 100 amino acids. A, Left shows the distribution of numbers of N-glycosylation sequence motifs (N-X-S/T) per 100 amino acids. Proteins with more than five motifs were excluded from this graph. A, Center shows the distribution of numbers of C-glycosylation sites per 100 amino acids predicted by using GlycoMine. Proteins with more than four predicted sites were excluded. A, Right shows the distribution of numbers of cysteine residues per 100 amino acids. Proteins with >10 cysteines were excluded. (B) Distribution of the number of ɑ-helical TM domains per protein. The pie chart shows the proportion of GPCRs within the set of surface proteins with seven TM domains (red bar). (C) Distribution of the length of ɑ-helical TM domains. (D) Classification of surface and nonsurface proteins into functional classes. *P < 10−2. Functional classes are numbered as follows: 1, GPCRs; 2, receptor-type tyrosine kinases; 3, receptors of the Ig superfamily; 4, scavenger receptors; 5, other receptors; 6, channels; 7, solute carrier superfamily; 8, active transporters; 9, auxiliary transport proteins; 10, other transporters; 11, oxidoreductases; 12, transferases; 13, hydrolases; 14, lyases; 15, isomerases; 16, ligases; 17, structure/adhesion proteins; 18, ligand proteins; and 19, proteins of unknown function. (E) Overlap of proteins of the human surfaceome annotated in UniProt, predicted by YLoc (13), predicted by da Cunha et al. (17), and predicted by SURFY. (F) Protter image of human MEGF9. N-X-S/T motifs are marked in light blue, with the corresponding asparagine (N) in dark blue. CSC identified peptides are marked in purple. (G) Half-life distributions of surfaceome proteins, transcription factors (TF), and all quantified proteins from Beck et al. (65). Misc., miscellaneous.
Figure 3
Surfaceome expression in 610 cancer cell lines. (A) Distribution of cell-specific surfaceome diversity (count of expressed surfaceome genes; left axis), sorted from large to small. Cell lines are colored based on their tissue type, as indicated by the color code. The straight gray line marks the average, and the dashed line marks the median surfaceome diversity. The sum of the expressed surfaceome genes is indicated by the black line corresponding to the right axis. (B) Distribution of surfaceome diversities (count of expressed surfaceome genes) based on tissue type. Tissues are color-coded as in A. The number of cell lines belonging to each tissue is indicated on the horizontal axis. (C) Scatter plot of count of expressed surfaceome genes vs. physical cell size. Squared Pearson correlation coefficient is indicated. (D) Box plots of the surfaceome gene expression level distribution for each cell line, sorted based on surfaceome diversity as in A from large to small. The black range represents the interquartile range; whiskers are depicted in gray. (E) Distribution of log2 expression level of PD-L1 cell lines with the highest expression are indicated. (F) Surfaceome genes sorted by number of cell lines in which each gene is expressed enabled categorization into five groups. Functional classification for each group of genes based on Almén et al. (1) is shown in the bar chart in F, Inset. Misc., miscellaneous.
Figure 4
Voronoi tree maps generated on wlab.ethz.ch/surfaceome. Maps for RAMOS (A), HT-29 (B), and IMR-32 (C) are shown. RPKME values of each cell line were scaled from 0 to 1 and mapped onto the whole in silico surfaceome. Light color indicates low expression; dark color indicates strong expression. White genes are not expressed. Characteristic functional protein groups of these cell lines are highlighted on the
Figure 5
Surfaceome changes during neurogenesis. (A) Left axis: Surfaceome gene level distribution from day 0 to day 22. Right axis: The red line shows the total number of expressed surfaceome genes, and the brown line shows the sum of expression levels over all expressed surfaceome genes. Transcriptomic data and definition of developmental stages (1, pluripotency stage; 2, differentiation initiation stage; 3, neural commitment stage; 4, NPC proliferation stage; 5, neuronal differentiation stage) were obtained from Li et al. (52). (B) Expression of selected gap junction genes from day 0 to day 22. (C) Identified clusters among surfaceome gene expression profiles based on c-means soft clustering. Red, higher correlation with cluster; light blue, lower correlation with cluster. (D) Voronoi tree map of log2 expression ratios between day 0 and day 22; the darker means more expressed at day 0, and the brighter color means more expressed at day 22. Surfaceome genes are hierarchically grouped by functional classification [receptors (orange), transporters (purple), hydrolases (dark blue), unclassified (blue), and miscellaneous (green)].
Welcome to the in silico human surfaceome –
Despite the fundamental importance of the surfaceome as a signaling gateway to the cellular microenvironment, it remains difficult to determine which proteoforms reside in the plasma membrane and how they interact to enable context-dependent signaling functions. We applied a machine learning approach utilizing domain-specific features to develop the accurate surfaceome predictor SURFY and used it to define the human in silico surfaceome of 2,886 proteins. The in silico surfaceome is a new public biomedical resource which can be used to filter multi-omics data to uncover cellular phenotypes and new surfaceome markers.
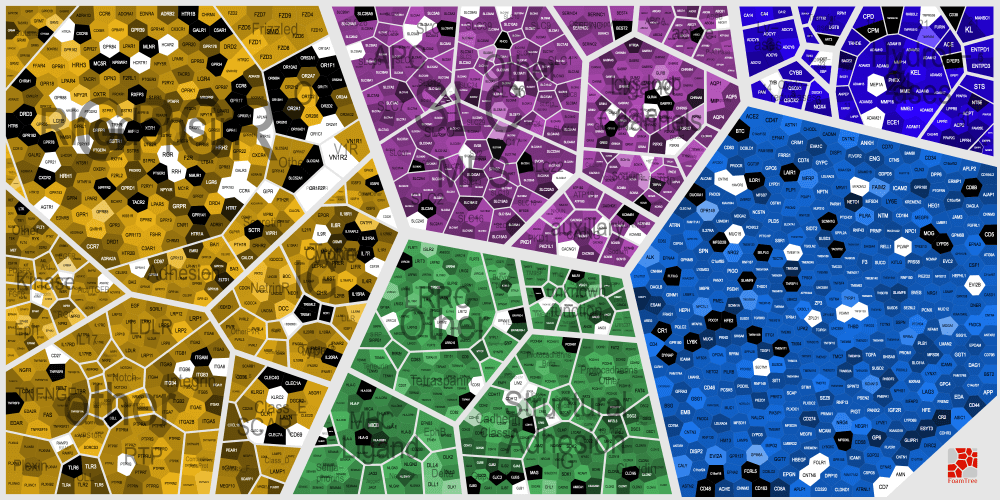
Surfaceome Adaptation during Neurogenesis
Map of log2 expression ratios in day 0 (black) vs. day 22 (white) in a transcriptional dataset of human embryonic stem cells (hESCs) and neural progenitors at various stages of differentiation (Li et al. J Biol Chem, 2017)
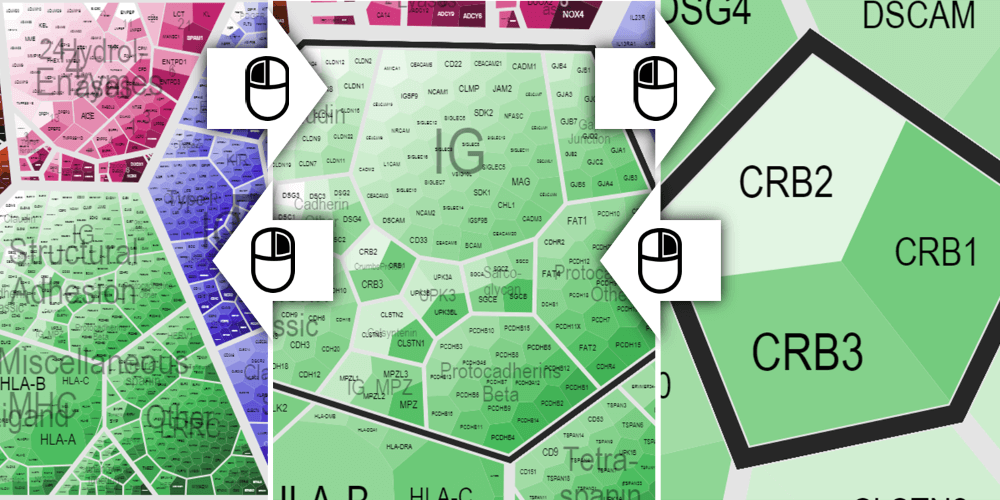
Functional Classification Navigation
Use you mouse to navigate the hierarchical functional classification visualization. A left-click selects and zooms in to a subclass, while and a right-click selects and zooms out to the class up in the hierarchy. The table will be filtered according to the selected functional (sub-)class. Alternatively you can also select a (sub-)class from the list in the upper-left corner.
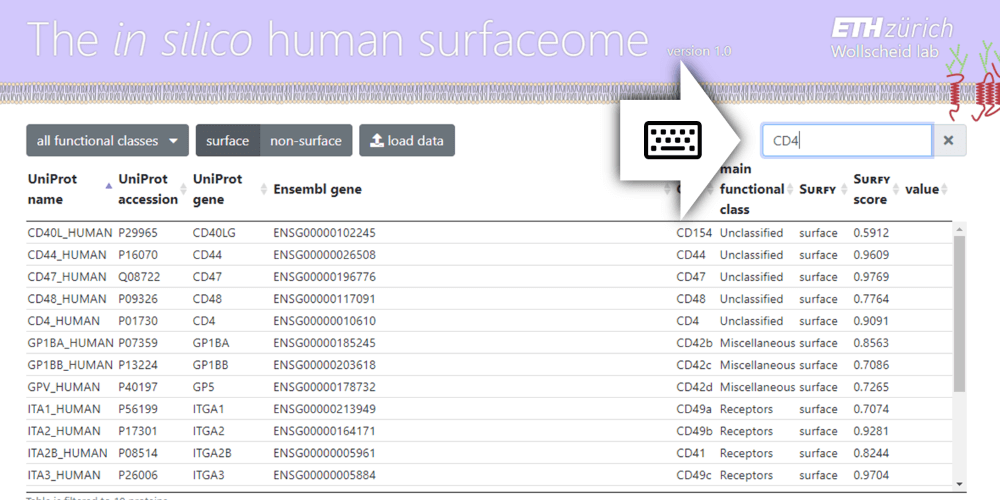
Find your Protein
To find your favorite gene/protein, use the search box in the upper right corner. The protein table will be filtered to show your search results. Clicking on a row in the table will highlight the corresponding gene in the functional classification map.
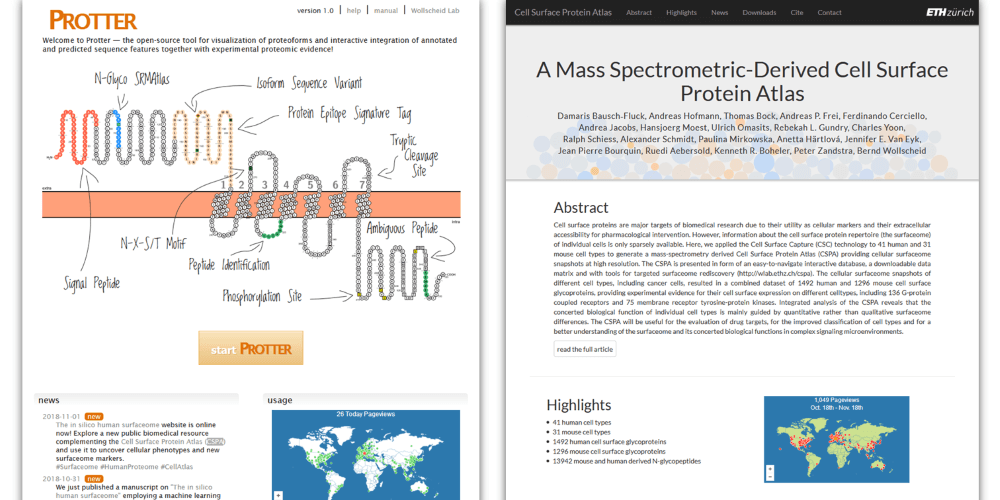
More Surfaceome Resources provided by the Wollscheid Group
- Protter, an open-source tool specially helpful for visualization of cell surface proteins.
- CSPA, a mass spectrometric-derived Cell Surface Protein Atlas based on 41 human cell types and 31 mouse cell types.
news
- 2018-11-20 new
- The surfaceome paper was recommended by a F1000 member of being of special significance in its field!

- 2018-10-31 new
- We just published a manuscript on "The in silico human surfaceome" employing a machine learning approach (SURFY). This new public biomedical resource can be used to uncover cellular phenotypes and new surfaceome markers!
#Surfaceome #HumanProteome #CellAtlas #PNAS - 2018-10-10 new
- Our review/perspective on "Surfaceome Nanoscale Organization and Extracellular Interaction Networks" in "Current Opinion in Chemical Biology" is now online! Here's a share link for free access for 50 days: review
Thanks to editors Ileana Cristea & @lilley_ks!
links
- Protter
- An open-source tool specially helpful for visualization of cell surface proteins. It includes proteoforms and interactive integration of annotated and predicted sequence features together with experimental proteomic evidence!
- Cell Surface Protein Atlas
- A Mass Spectrometric-Derived Cell Surface Protein Atlas based on in depth analysis of 41 human cell types and 31 mouse cell types.
usage
how to cite
The in silico human surfaceome has been published in PNAS:
The in silico human surfaceome.
Bausch-Fluck D, Goldmann U, Müller S, van Oostrum M, Müller M, Schubert OT, Wollscheid B.
Proc Natl Acad Sci U S A. 2018 Nov 13;115(46):E10988-E10997.