How many people/proteins/proteoforms reside in this space ("Surfaceome")? Similar to humans, proteins don't act alone. Function is encoded in dynamic protein-protein interactions. How are these proteoforms organized in signaling islands/networks in order to fulfill specific cellular functions ("Interactome")? What are the ligands interacting with the surfaceome to communicate information from other cells & tissues in the body? What goes wrong in these signaling islands if we get sick? Check out the baseball fields: A baseball field in the park would be one of those functional interaction islands upon which proteins – the players – group themselves for a particular cellular function – in this case, the baseball game. If the players leave the field, they lose their ability to function in this way. It's all about baseball & functional nanoscale organization of the surfaceome!
"Surfaceome nanoscale organization and extracellular interaction networks"The goal of the Wollscheid laboratory is to functionally understand the cellular surfaceome and its signaling islands as a complex information gateway connecting the intracellular to the extracellular interactome. We develop and apply next generation technologies at the interface of biology, chemistry, medicine and bioinformatics to generate unprecedented data to establish the surfaceome proteotype and its signaling interaction network. This digital proteotype data layer provides the basis for generating qualitative and quantitative surfaceome models explaining how molecular nanoscale organization influences cellular signaling and biological function.
Based on a specific biomedical question we measure millions of spectra derived from biospecimen using mass spectrometry.
Using complex bioinformatic pipelines & statistical models we integrate the data and establish the proteotype of a biospecimen.
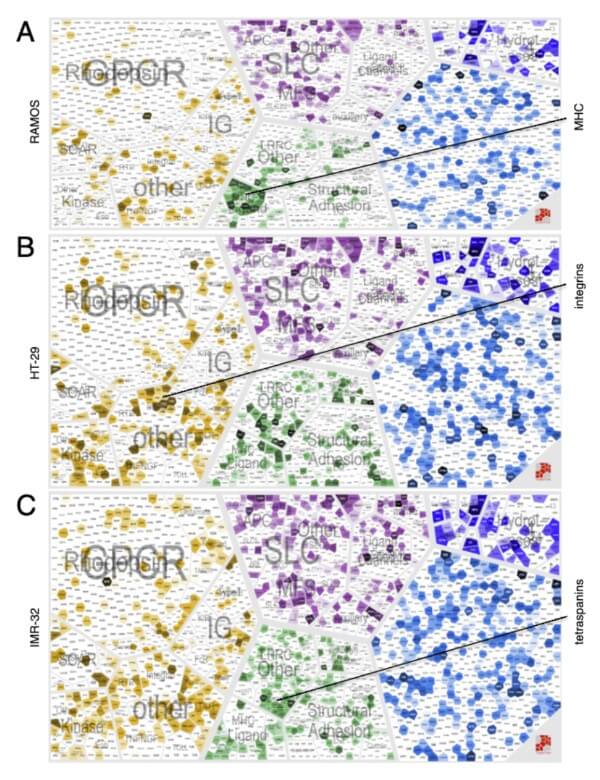
We analyze the spatiotemporal dynamics of the biological system/proteotype under perturbation and generate hypotheses in order to explain the phenotype and molecular processes involved.
Using a variety of Systems Biology technologies we validate our findings and turn raw data into new biological insights & basic knowledge on how cells communicate.
The Wollscheid lab developed a set of key Technologies & Resources which not only enable quantitative surfaceome discovery, but also the elucidation of functional protein-protein interactions. Life depends on interactions. The same holds true on the molecular level. Understanding molecular interaction networks is a prerequisite to understand basic biology and in turn, human health.
Surfaceome Screening
Cell Surface Capture (CSC) technology enables qualitative & quantitative surfaceome discovery in order to phenotype cells without antibodies.
Learn MoreSurfaceome Interactome Screening
TRICEPS/HATRIC-based Ligand Receptor Capture (LRC) technology enables the identification of receptors for orphan ligands on living cells in order to deconvolute complex ligand receptor interactions. Technology patented by ETH licensed to dual systems.
MoreIntercellular Surfaceome Screening
LUX-MS enables the light-mediated proximity detection of cell surface signaling island & neighborhoods on living cells.
Learn MoreSurfaceome KnowledgeBase
The Cell Surface Protein Atlas (CSPA) is a new public resource of experimentally verified cell surface accessible markers and targets for therapeutics.Check out the CSPA website.
Learn MoreSurfaceome Visualization
PROTTER is an open source tool for the visualization of proteoforms and interactive integration of annotated and predicted sequence features together with experimental proteomic evidence. Check out the PROTTER website.
Learn MoreSurfaceome Data Analysis
The in silico human surfaceome is based on the machine learning algorithm SURFY and represents a new public resource for the visual interrogation of multi-omics datasets for the biomedical community. Check out the SURFY website.
Learn More
We are always looking for highly motivated people to join our team!
Get in touch if you are
interested.
Still not clear who we are and what we do? Or do you have other questions? Perhaps your questions are answered below, but don't hesitate to contact us if they're not.
We are a publicly funded laboratory headed by Prof. Dr. Bernd Wollscheid at the Institute of Translational Medicine and the Department of Health Sciences and Technology of the Swiss Federal Institute of Technology / ETH Zurich in Switzerland.
Our research is supported by public funding such as


We are always looking for enthusiastic people to join our team. We currently have several vacancies that we are looking to fill. Please apply by e-mailing your CV and an explanation of your motivation to Bernd Wollscheid.